Translatomics for viral infection, cytokine responses, and plasma cell differentiation
Recent Publications Harnessing the Power of Translatomics
Every week we provide a digest of a small number of recent interesting papers in the field of translatomics.
In this week’s Sunday papers,
- Kerkhofs et al. find that respiratory syncytial virus (RSV) infection bypasses the typical stress response, boosts polysome loading, and shifts translation toward long, AU-rich transcripts—including viral RNAs—while the translation efficiency in highly translated host mRNAs is reduced.
- Liu et al. find that cytokine release syndrome (CRS) during CAR T therapy is driven by widespread translational activation in monocytes and macrophages, with BCAP–PI3K–RSK–EIF4B signalling enhancing IL-6 translation, and that targeting RSK reduces protein synthesis and alleviates CRS symptoms.
- Chen et al. find that the RNA-binding proteins Csde1 and Strap cooperatively regulate translation and mRNA stability to control plasma cell differentiation, promoting Bach2 mRNA decay and enhancing Atp2a2 translation, with their physical interaction being essential for function.
Respiratory syncytial virus (RSV) enhances translation of virus-resembling AU-rich host transcripts
Virology Journal, 2025
Kerkhofs, K., Guydosh, N.R. and Bayfield, M.A.
Viral infection often reshapes the host’s translational landscape as viruses compete with cellular mRNAs for ribosomes. Many cellular stresses suppress 5′-cap-dependent translation through phosphorylation of eIF2α, which blocks translation initiation and promotes stress granule formation. Respiratory syncytial virus (RSV), a negative-sense RNA virus, produces capped and polyadenylated transcripts that closely resemble host mRNAs and are translated by host ribosomes in the cytoplasm. Although viral infection typically activates the PKR kinase and triggers eIF2α phosphorylation, RSV counteracts this by using its nucleocapsid protein (N) to bind PKR and prevent inhibition of translation. However, reports on stress granule formation during RSV infection remain conflicting, and it is unclear how RSV efficiently competes with host transcripts for the translational machinery. The authors therefore sought to investigate this further.
They first examined whether RSV infection activates the integrated stress response. Western blotting showed only minimal phosphorylation of eIF2α compared with arsenite-treated cells, and immunofluorescence using stress-granule markers (PABP, G3BP) confirmed that RSV infection does not trigger stress-granule formation. Polysome profiling revealed an unexpected increase in polysome occupancy in infected cells, indicating enhanced ribosome loading onto mRNAs rather than global translational repression. To determine which transcripts were affected, the authors combined polysome fractionation with RNA-seq (polysome-seq) to quantify total and ribosome-associated mRNAs and calculate translation efficiency (TE). Many changes in polysome association reflected transcriptional changes, such as upregulation of interferon-stimulated genes, while others were driven by altered TE. Notably, transcripts with low baseline TE had increased ribosome loading, whereas highly translated transcripts lost relative efficiency. Re-analysis of published VSV (vesicular stomatitis virus, another negative-sense single-stranded RNA virus), infection datasets revealed a similar redistribution of ribosomes. Sequence analysis showed that longer, AU-rich transcripts—including viral RNAs—were preferentially enriched in heavy polysomes, suggesting that infection biases translation toward this class of mRNAs.
Learn more about EIRNABio’s polysome profiling services here.
Discovery, delineation, and therapeutic targeting of a hyper-translation pathway driving cytokine release syndrome
Cell Reports Medicine, 2026
Liu, X., Li, J., Zhang, Y., Liu, Y., Wang, C., Wang, Y., Liu, Y., Yang, Y., Su, Y., Lu, Y., Wang, W, Fu, Y.-X., Lin, X., Pan, D., Han, W., and Hu, X.
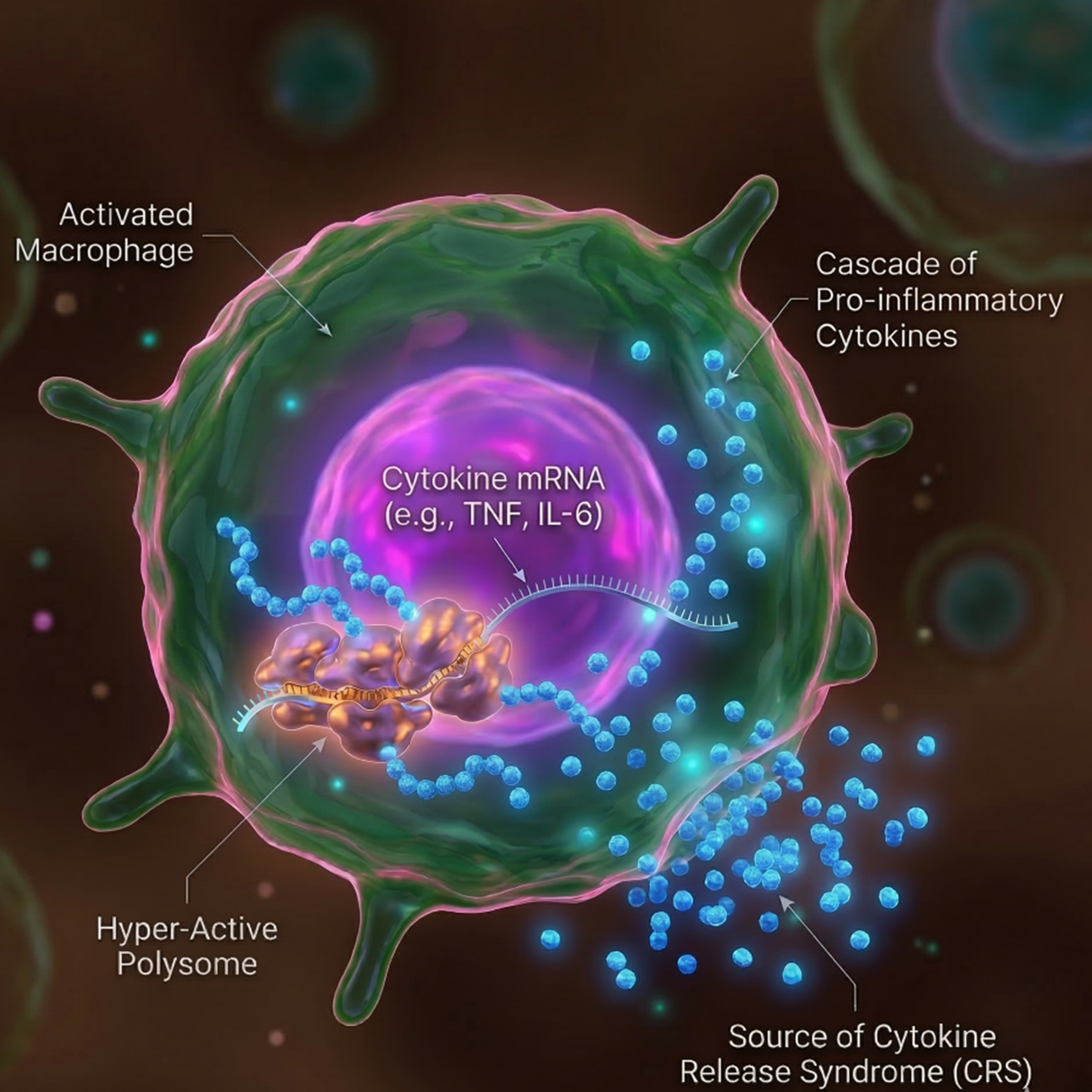
Inflammation is a tightly regulated process essential for development, immunity, and tissue homeostasis. While normally self-limiting, dysregulated inflammation can lead to disease. One severe example is cytokine release syndrome (CRS), a potentially life-threatening complication frequently observed in patients receiving CAR T cell therapy. CRS is characterized by excessive production of pro-inflammatory cytokines, particularly IL-6, largely driven by monocytes and macrophages. Although inflammatory responses have traditionally been studied at the level of transcription, the role of translational regulation remains less explored. Notably, the β1-blocker metoprolol was recently found to reduce IL-6 production in CRS without affecting IL-6 mRNA levels, instead lowering its translation efficiency. These observations suggest that translational control may be a key regulator of pathological inflammation, prompting the authors to investigate this further.
To characterize cytokine release syndrome (CRS) in patients receiving CAR T therapy, the authors analysed PBMCs collected before and during CRS. Quantitative proteomics by mass spectrometry revealed strong upregulation of proteins involved in translation, including ribosomal proteins and initiation factors. SUnSET assays measuring puromycin incorporation further showed that CD14⁺ monocytes exhibit markedly increased protein synthesis during CRS, indicating widespread translational activation. Using an in vitro CAR T killing supernatant (KS) model, the authors confirmed this phenotype and performed ribosome profiling (Ribo-seq), which demonstrated increased translation efficiency (TE) across roughly one-third of protein-coding genes. Parallel RiboTag-seq translatome profiling in macrophages showed that most CRS-associated changes occur at the level of translation rather than transcription. A CRISPR screen in primary macrophages using IL-6 translation as a readout identified the signalling adaptor BCAP (PIK3AP1) as a key regulator. Genetic deletion, polysome profiling, and Ribo-seq showed that BCAP drives macrophage hyper-translation via a PI3K–RSK–EIF4B signalling axis. Finally, genetic ablation or pharmacological inhibition of RSK reduced IL-6 translation and global protein synthesis without affecting mRNA levels. RSK inhibition also alleviated CRS symptoms in humanized mouse models, highlighting translational control as a potential therapeutic target.
Learn more about EIRNABio’s ribosome profiling services here.
A Csde1-Strap complex regulates plasma cell differentiation by coupling mRNA translation and decay
Nature Communications, 2025
Chen, P., Lin, L., Lin, X., Liao, K., Qiang, J., Wang, Z., Wu, J., Li, Y., Yang, L., Yao, N., Song, H., Hong, Y., Liu, W.-H., Zhang, Y., Chang, X., Du, D., and Xiao, C.
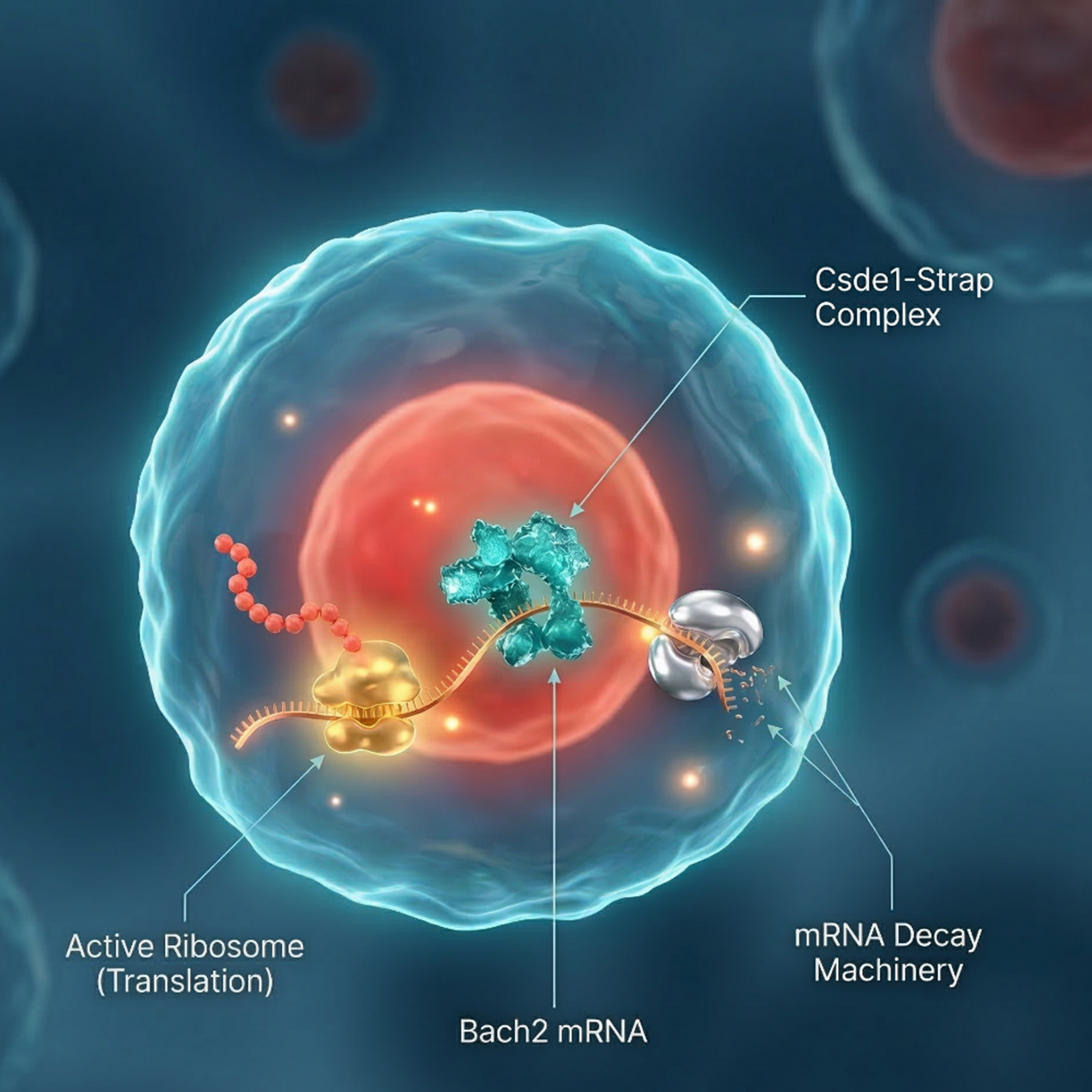
Precise control of protein synthesis is essential for dynamic cellular processes such as activation, proliferation, and differentiation. While transcription, epigenetic regulation, and protein degradation are well-established regulators of gene expression, the roles of mRNA translation and decay remain less well understood. Increasing evidence suggests that many mRNAs are degraded co-translationally and that regulatory elements within untranslated regions and coding sequences coordinate translation with mRNA stability. B cells provide an ideal model to study these mechanisms because they undergo tightly regulated activation and differentiation during immune responses, ultimately forming plasma and memory cells. Although transcription factor networks controlling plasma cell differentiation are well characterized, emerging evidence indicates that RNA-binding proteins and microRNAs also play key regulatory roles. The authors therefore sought to investigate this further.
To identify RNA-binding proteins (RBPs) involved in plasma cell differentiation, the authors performed mRNA interactome capture (RIC) coupled with mass spectrometry across naïve B cells, germinal center–like B cells (iGCB), and plasma cells (iPC). This revealed hundreds of RBPs and highlighted strong enrichment of translation-related RBPs during B cell activation. A CRISPR–Cas9 pooled screen targeting 759 RBPs then identified regulators of B cell expansion and plasma cell differentiation, with many hits linked to translational control. Among these, Csde1 and Strap emerged as key factors. CRISPR knockouts and flow cytometry assays showed that loss of either protein impaired plasma cell differentiation, while conditional knockout mice and immunization experiments demonstrated reduced germinal center responses and antigen-specific antibody production in vivo. To uncover their targets, the authors used eCLIP-seq, showing that Csde1 and Strap bind largely overlapping sets of mRNAs, predominantly within coding regions. Quantitative proteomics, RNA-seq, and ribosome profiling revealed that these proteins cooperatively regulate translation and stability of shared targets. Mechanistically, Csde1–Strap promotes translation-induced decay of Bach2 mRNA and enhances translation of Atp2a2, both of which are critical for plasma cell differentiation. Finally, mutational and rescue experiments demonstrated that physical interaction between Csde1 and Strap is essential for RNA binding and their regulatory function.
Learn more about EIRNABio’s ribosome profiling services here.



