
Powered by Ribo96: High-Resolution Translation Analysis at Unmatched Scale & Cost
Understanding gene expression at the level of translation is critical for unlocking the full potential of early-stage drug discovery, functional genomics, and personalised medicine. Yet traditional transcriptomic and proteomic approaches fall short, with transcriptomics hinting at what might be expressed and proteomics shows what has been produced. This gap leads to misinterpretations, missed targets, and delays in therapeutic development.
Bridging the divide between RNA-Seq and Proteomics
Ribosome profiling (Ribo-seq) bridges this divide by offering direct, high-resolution insights into real-time protein synthesis. And at EIRNABio, our ribosome profiling services are powered by Ribo96 – our proprietary, high-throughput platform enabling the simultaneous analysis of up to 96 samples in a single run. With this technology, we combine scientific precision with industrial-scale capacity, giving you rapid, robust, and reproducible results across diverse biological conditions – all done at a fraction of the cost associated with traditional ribosome profiling methodology


What is Ribosome Profiling?
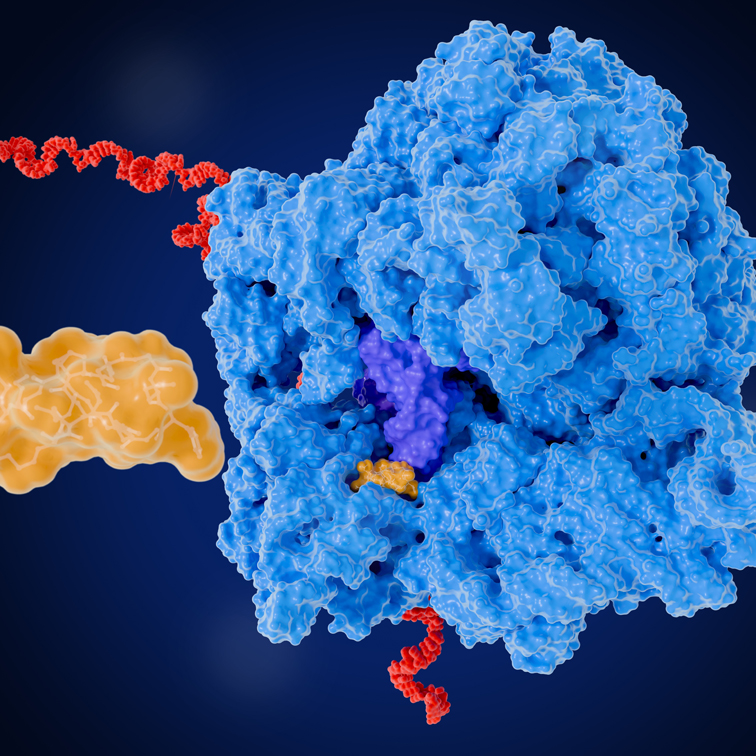
Ribosome profiling is a high-throughput sequencing technique that empowers researchers to investigate translation with single nucleotide resolution. The unprotected mRNA is degraded by nuclease digestion, leaving ribosome-protected fragments (RPFs), which are then isolated and sequenced. By mapping these captured sequences to annotated transcripts, ribosome profiling captures a comprehensive picture of which genes are being actively translated and to what extent, providing invaluable data on gene expression regulation, translational efficiency, and codon usage.
Protocol Overview: Cells are first lysed under conditions that maintain the integrity of ribosome-mRNA complexes. Translating ribosomes are then pharmacologically stalled, using translation inhibitors such as cycloheximide, preserving their position on transcripts. Lysates subsequently undergo nuclease digestion to remove any unprotected mRNA, while RPFs are protected from digestion.
The resulting RPFs, which are approximately 28-32nt in length, are purified and size selected on Urea PAGE gels. Contaminating fragments derived from rRNA are depleted from the sample before the enriched RPFs are converted into a cDNA library, and subjected to deep sequencing for high-resolution analysis of translation.
Key Features of Ribosome Profiling
Quantitative Measurement
Provides quantitative data on ribosome occupancy on mRNA, offering insights into translational efficiency and protein synthesis across transcripts
Detailed Resolution
Allows for a fine-grained understanding of codon usage, revealing variations in translation efficiency across different conditions.
Dynamic Insights
Offers a time-resolved perspective on translation, enabling the study of how protein synthesis responds to experimental conditions, such as environmental changes, gene editing, or therapeutic interventions.
Broad Applicability
Useful across a variety of research areas, from cancer biology to neuroscience, facilitating investigations into disease mechanisms and cellular responses to drug treatment. Ribosome profiling offers insights for novel therapeutic interventions, such as gene therapy and ATMPs, while also having applications for biomarker discovery through exploration of the hidden proteome.
Ribo96: The Engine Behind EIRNABio’s Ribosome Profiling
Traditional ribosome profiling is labour-intensive, costly, and prone to variability. Our Ribo96 technology changes that. Ribo96 is EIRNABio’s proprietary high-throughput platform that miniaturises and automates ribosome profiling in a 96-well format. By streamlining complex wet-lab steps and integrating rigorous quality control, Ribo96 enables scalable and reproducible profiling across conditions, replicates, or treatments—all in a single sequencing-ready batch.
Key Advantages of Ribosome Profiling
High-Throughput
Profile up to 96 samples simultaneously—ideal for drug screens, time-course experiments, or cohort analyses
Cost-Efficient
Lower per-sample costs compared to conventional Ribo-seq methods
Reproducible
Automated workflows reduce batch effects and variability
Quality-Driven
Each run is rigorously assessed for key metrics like triplet periodicity and footprint distribution
Challenges Associated with ‘traditional’ Ribosome Profiling
Data Generation
Despite the notable insights that ribosome profiling provides, a number of barriers exist which limit its widespread implementation.
Challenge 1
Technically Challenging: Generating high-quality Ribo-seq data demands broad molecular biology expertise. Key steps like lysis, digestion, and footprint selection require precision, with decades of experience helping to avoid common pitfalls.
Challenge 2
Steep & Expensive Learning Curve: Implementing ribosome profiling in-house requires major investment in equipment and staff training. The protocol is complex and labor-intensive, with a high risk of costly trial-and-error for inexperienced teams.
Challenge 3
Poor Data Quality leading to repeat experiments: Ribo-seq protocols take 5–10 days, but sample issues can yield poor data. Identifying these problems often takes months, requiring researchers to repeat the entire process from library prep to analysis.
Solution 1
Expertise: Led by translatomics pioneers, we tailor each project to meet exact requirements, delivering high-quality Ribo-seq data across sample types. Explore key publications from our founders that shaped the field.
Solution 2
Experienced Reliable Providers: With decades of experience and hundreds of completed projects, we offer a reliable, cost-effective alternative to in-house setup. Clients trust us to deliver consistently high-quality ribosome profiling data. See a selection of customer publication.
Solution 3
Excellence: We apply early quality control to ensure only high-quality Ribo-seq libraries advance to sequencing. Key metrics like triplet periodicity—critical for interpreting translation accuracy—are rigorously assessed. Learn more about quality features.
Challenge 2
Steep & Expensive Learning Curve: Implementing ribosome profiling in-house requires major investment in equipment and staff training. The protocol is complex and labor-intensive, with a high risk of costly trial-and-error for inexperienced teams.
Solution 2
Experienced Reliable Providers: With decades of experience and hundreds of completed projects, we offer a reliable, cost-effective alternative to in-house setup. Clients trust us to deliver consistently high-quality ribosome profiling data. See a selection of customer publication.
Challenge 3
Poor Data Quality leading to repeat experiments: Ribo-seq protocols take 5–10 days, but sample issues can yield poor data. Identifying these problems often takes months, requiring researchers to repeat the entire process from library prep to analysis.
Solution 3
Excellence: We apply early quality control to ensure only high-quality Ribo-seq libraries advance to sequencing. Key metrics like triplet periodicity—critical for interpreting translation accuracy—are rigorously assessed. Learn more about quality features.
Data Analysis
Challenge 1
The positional information that ribosome profiling captures is a vast resource that enables investigation of an extensive variety of translational phenomena, including translational efficiency, ribosome stalling, uORF-mediated regulation, stop codon readthrough, ribosomal frameshifting, and exploration of the dark proteome for microprotein identification. The wealth of applications to which this rich dataset lends itself can be overwhelming to newcomers looking to gain biological insights specific to their interests.
Challenge 1
Complexity of Data: Ribosome profiling data offers vast insights into translation, from efficiency to microprotein discovery. However, its complexity can overwhelm newcomers seeking specific biological interpretations within this rich dataset.
Challenge 2
Limitations of existing ribosome profiling analysis tools: Ribosome profiling analyses often require multiple tools, each targeting specific translational phenomena. Bioinformaticians must invest significant time and effort benchmarking and running separate analyses to fully interpret the data.
Challenge 3
Static Outputs: Ribosome profiling’s nucleotide-level detail is often underused because many tools provide only static plots and spreadsheets, making deeper insights hard to uncover without intensive manual analysis.
Challenge 4
Avoiding Errors in Codon Decoding Analysis: Accurate ribosome profiling depends on correct A-site inference. Misassigning the codon decoded by ribosome footprints can lead to misleading or incorrect biological conclusions.
Solution 1
Introducing EIRNABio Connect, the interactive browser based all-in-one bioinformatics platform for the analysis of ribosome profiling data. Designed to carry out most of the “heavy lifting” for experienced bioinformaticians, whilst at the same time empowering those without in-depth bioinformatics expertise to dynamically interact with and interrogate their data.
Solution 1
Easy-to-use Analysis Platform: EIRNABio Connect is an interactive, browser-based platform that simplifies ribosome profiling analysis, handling complex tasks for experts while enabling non-experts to explore and interpret their data dynamically without deep bioinformatics skills.
Solution 2
Interactive Dynamic Visualization: EIRNABio Connect offers interactive visualizations, allowing users to click data points, zoom in, and explore ribosome profiles at codon-level resolution, unlocking the full potential of nucleotide-level ribosome profiling data.
Solution 3
Comprehensive Functionality: EIRNABio Connect offers a comprehensive suite of diverse analytical modules, including gene expression, ribosome distribution, pause prediction, codon occupancy, and alternative ORF detection—all integrated for streamlined, in-depth ribosome profiling analysis.
Solution 4
Comprehensive Support: EIRNABio handles all data processing before platform upload and offers help pages, videos, workshops, and consultations to ensure customers accurately interpret their ribosome profiling data and maximize insights.
Challenge 1
Complexity of Data: Ribosome profiling data offers vast insights into translation, from efficiency to microprotein discovery. However, its complexity can overwhelm newcomers seeking specific biological interpretations within this rich dataset.
Solution 1
Easy-to-use Analysis Platform: EIRNABio Connect is an interactive, browser-based platform that simplifies ribosome profiling analysis, handling complex tasks for experts while enabling non-experts to explore and interpret their data dynamically without deep bioinformatics skills.
Challenge 2
Limitations of existing ribosome profiling analysis tools: Ribosome profiling analyses often require multiple tools, each targeting specific translational phenomena. Bioinformaticians must invest significant time and effort benchmarking and running separate analyses to fully interpret the data.
Solution 2
Comprehensive Functionality: EIRNABio Connect offers a comprehensive suite of diverse analytical modules, including gene expression, ribosome distribution, pause prediction, codon occupancy, and alternative ORF detection—all integrated for streamlined, in-depth ribosome profiling analysis.
Challenge 3
Static Outputs: Ribosome profiling’s nucleotide-level detail is often underused because many tools provide only static plots and spreadsheets, making deeper insights hard to uncover without intensive manual analysis.
Solution 3
Interactive Dynamic Visualization: EIRNABio Connect offers interactive visualizations, allowing users to click data points, zoom in, and explore ribosome profiles at codon-level resolution, unlocking the full potential of nucleotide-level ribosome profiling data.
Challenge 4
Avoiding Errors in Codon Decoding Analysis: Accurate ribosome profiling depends on correct A-site inference. Misassigning the codon decoded by ribosome footprints can lead to misleading or incorrect biological conclusions.
Solution 4
Comprehensive Support: EIRNABio handles all data processing before platform upload and offers help pages, videos, workshops, and consultations to ensure customers accurately interpret their ribosome profiling data and maximize insights.


Application Areas
Complete Gene Expression
Complete gene expression analysis with EIRNA Bio’s ribosome profiling service offers unparalleled insights into protein synthesis by capturing the actively translating mRNAs in cells.
Learn More
Global Translation Rates
Ribosome profiling for Global Translation Rates at EIRNA Bio enables researchers to quantify protein synthesis across the entire cellular transcriptome, providing dynamic, nucleotide-level insights into translational activity.
Codon Optimality
EIRNABio’s ribosome profiling for Codon Optimality enables researchers to study how codon usage impacts translational efficiency, revealing patterns in protein synthesis that traditional transcriptomics overlook.
Off-Target Expression Effects
Ribosome profiling at EIRNABio provides researchers with a powerful method to assess off-target expression effects across the transcriptome.
uORF Regulatory Activity
EIRNABio’s ribosome profiling enables in-depth analysis of upstream open reading frames (uORFs), which regulate gene expression by modulating translation.
Ribosome Pausing
Ribosome pausing analysis using EIRNABio’s ribosome profiling service enables researchers to explore translation regulation and quality control mechanisms at codon resolution.
Nonsense Suppression/
Readthrough
EIRNABio’s ribosome profiling for nonsense suppression and stop codon readthrough provides precise insights into translation termination failures.
Micropeptide Prediction
Micropeptides are small proteins often overlooked in traditional gene annotation methods. EIRNA Bio’s ribosome profiling service provides a high-resolution approach to identify micropeptide-coding regions, revealing hidden elements of the proteome.
Translation Start Sites
EIRNABio’s ribosome profiling facilitates the identification of translation start sites with single-nucleotide resolution. By capturing ribosome positions at initiation codons, this service distinguishes canonical (AUG) and non-canonical start sites, enabling researchers to explore alternative initiation mechanisms.
Learn More
Biomarker Discovery
Biomarkers are crucial for disease diagnosis, monitoring, and therapeutic development. EIRNABio’s ribosome profiling service enables the discovery of translational biomarkers by identifying differentially expressed proteins and translation dynamics with codon-level precision.
Personalized Medicine
Personalised medicine aims to tailor treatments based on individual genetic and molecular profiles. EIRNA Bio’s ribosome profiling service enables researchers to identify translational differences among patients by capturing real-time protein synthesis.
Ribosome Profiling Services FAQ
What is ribosome profiling, and how does it work?
Ribosome profiling is a cutting-edge technique that provides a snapshot of active translation in cells by capturing ribosome-protected mRNA fragments. This method enables researchers to identify which genes are being actively translated and provides detailed insights into protein synthesis, gene expression regulation, and translation dynamics at the nucleotide level.
How is ribosome profiling different from other gene expression methods?
Traditional gene expression methods like RNA sequencing measure mRNA levels, but they do not reveal if the mRNA is actively being translated into proteins. Ribosome profiling goes beyond this by identifying the mRNAs bound to ribosomes, offering a direct view of active translation. This provides a higher resolution of cellular activity, especially in understanding how proteins are synthesised under different conditions.
What are the key applications of ribosome profiling?
Ribosome profiling has broad applications across various fields, including:
- Gene expression and regulation studies
- Translational control mechanisms
- Ribosome stalling and codon optimality
- Personalised medicine and biomarker discovery
- Investigating disease mechanisms in cancer, neuroscience, and more
What challenges are typically associated with ribosome profiling?
Some common challenges include:
- Technical complexity: Expertise in molecular biology is required for sample preparation and library construction.
- Cost: Setting up ribosome profiling internally can be expensive due to specialised equipment and training.
- Time: Standard protocols can take several days to complete, with the risk of poor-quality data requiring protocol repetition.
How does EIRNABio address these challenges?
EIRNABio’s ribosome profiling services are designed to overcome these challenges with:
- Expertise: Led by pioneers in the translatomics field, our team customises protocols to meet specific project needs, ensuring high-quality data.
- Reliability: We handle all aspects of the process, saving clients the time and expense of internal implementation.
- Early QC: We prioritise quality control to ensure that only high-quality libraries are sequenced, minimising the chance of failed experiments.
What is EIRNABio Connect, and how does it benefit my project?
EIRNABio Connect is our advanced, browser-based bioinformatics platform designed for ribosome profiling data analysis. It offers:
- Comprehensive analysis modules: Covering everything from differential gene expression to codon occupancy analysis.
- Interactive visualisation: Explore data down to the nucleotide level with dynamic plots and clickable data points.
- Extensive support: Our platform includes help resources, tutorials, and personal meetings to help you interpret your data accurately.
How long does a ribosome profiling project typically take?
Turnaround times for ribosome profiling projects vary depending on the sample type and experimental design. However, EIRNABio’s streamlined processes ensure fast, efficient delivery. We focus on early quality control to avoid delays caused by poor-quality data, allowing us to deliver high-quality results as efficiently as possible.
How can ribosome profiling contribute to drug discovery and personalised medicine?
Ribosome profiling provides real-time insights into translation and protein synthesis, making it invaluable in identifying novel therapeutic targets and biomarkers. It enables researchers to understand disease mechanisms at a translational level, accelerating drug discovery and supporting the development of personalised treatments based on individual genetic profiles.
Do you offer support for data interpretation?
Yes, we offer extensive support, including help pages, video demonstrations, and one-on-one consultations to assist in data interpretation. We can also provide in-person meetings and workshops tailored to your research goals.
What types of samples are compatible with ribosome profiling?
Ribosome profiling can be applied to a wide range of sample types, including cell lines, tissue samples, and even challenging sample types such as low-input or rare cell populations. We tailor our protocols to the specific needs of your project to ensure optimal results.
How do I get started with EIRNABio’s ribosome profiling services?
To get started, contact our team to discuss your project requirements. We offer a range of customisable service options and can help you design a ribosome profiling experiment that meets your specific research goals.
What is Ribo96, and how does it work?
Ribo96 is a high-throughput Ribosome Profiling service that maps active translation across 96 samples simultaneously by sequencing ribosome-protected mRNA fragments. This provides a detailed view of protein synthesis at nucleotide resolution.
How is Ribo96 different from traditional Ribosome Profiling?
While traditional Ribosome Profiling analyzes fewer samples per run, Ribo96 scales up to 96 samples, offering a cost-effective and efficient solution for large-scale studies without compromising data quality.
What are the key applications of Ribo96?
Ribo96 is ideal for drug screening, cohort studies, systems biology, personalized medicine, and exploring translational efficiency and regulation on a large scale.



Speak to an expert