Translatomics for microglia, mitochondria and stress granules
Recent Publications Harnessing the Power of Translatomics
Every week we provide a digest of a small number of recent interesting papers in the field of translatomics.
In this week’s Sunday papers,
- Flury et al. identify a maladaptive stress program in microglia that drives neurodegeneration.
- Dumas et al. investigate how RNA G-quadruplex (rG4) structures regulate translation of mRNAs that localize to mitochondria.
- Helton et al. show that loss of ribosome association is a prerequisite for mRNA entry into stress granules.
A neurodegenerative cellular stress response linked to dark microglia and toxic lipid secretion
Neuron, 2025.
Flury, A., Aljayousi, L., Park, H.J., Khakpour, M., Mechler, J., Aziz, S., McGrath, J.D., Deme, P., Sandberg, C., Ibáñez, F.G., Braniff, O., Ngo, T., Smith, S., Velez, M., Ramirez, D.M., Avnon-Klein, D., Murray, J.W., Liu, J., Parent, M., Mingote, S., Haughey, N.J., Werneburg, S., Tremblay, M.-E., and Ayata, P.
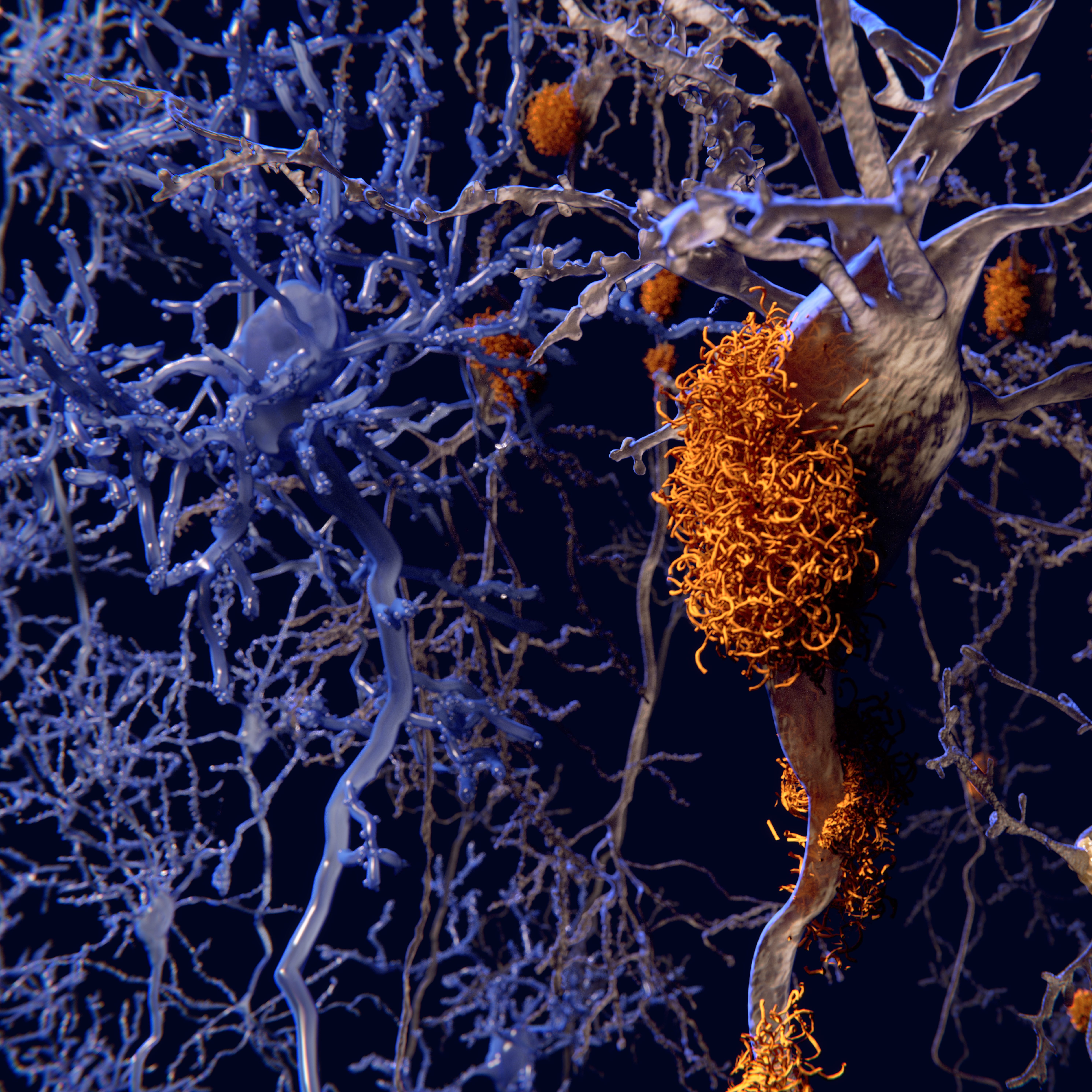
In this study the authors identify a microglial response triggered by chronic neuronal stress in Alzheimer’s disease (AD) models, characterized by integrated stress response (ISR) activation and metabolic/lipid remodelling. Central to this response is the emergence of dark microglia—a morphologically and functionally distinct microglial subtype associated with synaptic dysfunction and neuroinflammation. Using microglia-specific ribosomal profiling (TRAP), transcriptomics, lipidomics, and imaging approaches, the study shows that ISR-activated microglia activate pathways that promote the production and secretion of toxic lipid species, including long-chain phospholipids, sphingolipids, sterol esters, and glycerides. These lipids accumulate in the brain microenvironment and are closely associated with dark microglia activation.
Rather than providing neuroprotection, this stress response exacerbates neuronal damage, synapse loss, and inflammatory signalling. The study also reports that, despite ISR-associated eIF2α phosphorylation, microglia in disease contexts do not show sustained global translation arrest, consistent with chronic-stress adaptation. Microglia-specific ribosomal profiling further showed enrichment of ISR/stress and lipid metabolism programs, including upregulation of lipogenesis-associated genes such as Fasn. These molecular changes are consistent with the observed increase in toxic lipid secretion by microglia.
The findings suggest that neurodegeneration is not driven solely by protein aggregation or inflammation, but also by stress-induced lipid remodelling that actively reshapes the brain microenvironment. By showing that translational and metabolic shifts promote production of harmful lipids, the study highlights new therapeutic opportunities targeting microglial ISR signalling and de novo lipid synthesis which could help prevent synaptic loss and slow progression in disorders such as Alzheimer’s disease.
Learn more about EIRNABio’s ribosome profiling services here.
RNA G-quadruplexes control mitochondria-localized mRNA translation and energy metabolism
Nature Communications, 2025.
Dumas, L., Shin, S., Rigaud, Q., Cargnello, M., Hernández-Suárez, B., Herviou, P., Saint-Laurent, N., Leduc, M., Le Gall, M., Monchaud, D., Dassi, E., Cammas, A., and Millevoi, S.
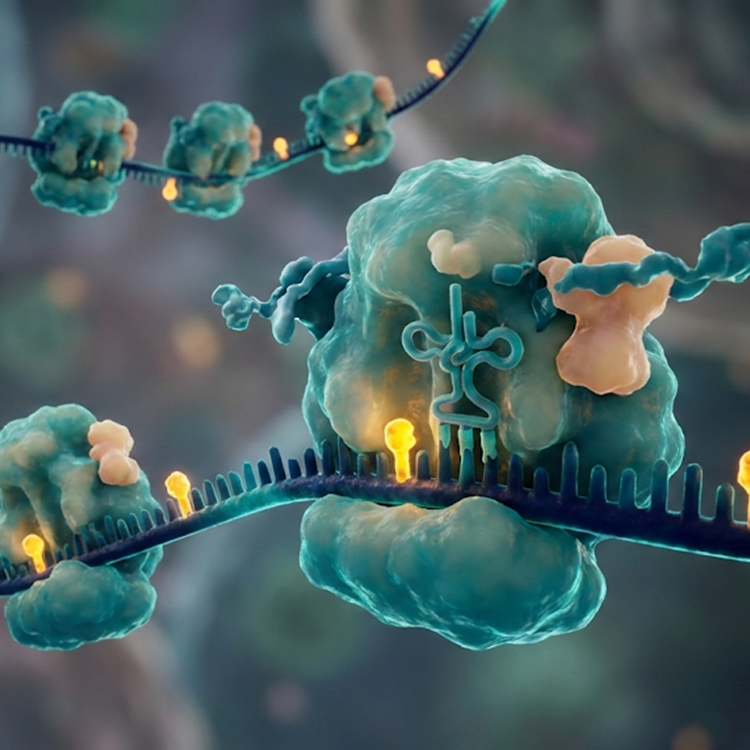
The study identifies RNA G-quadruplexes (rG4s) as critical regulators of mitochondrial function. It reveals that RG4 folding dynamics regulate translation, with hnRNP U as a key factor promoting translation of a subset of RG4-containing transcripts. A major finding is that many of the relevant RG4s are located in the 3′ UTRs of nuclear-encoded mitochondrial mRNAs. Disrupting RG4 dynamics, either by RG4-stabilizing ligands or by hnRNP U depletion, impairs translation of these transcripts, reduces mitochondrial protein expression, and disrupts Outer Mitochondrial Membrane localized translation. The key findings show that rG4s suppress the translation of mRNAs destined for the mitochondria, while the helicase DHX36 resolves these structures to promote protein synthesis. Disrupting rG4 dynamics impairs mitochondrial protein levels, leading to defective oxidative phosphorylation and shifted energy metabolism. The authors used polysome profiling coupled with RNA‑seq to assess how RG4s and their binding factor hnRNP U influence translation of nuclear‑encoded mitochondrial mRNAs. Polysome profiling revealed that depletion of hnRNP U significantly reduced the association of ~60 RG4‑containing nuclear‑encoded mitochondrial transcripts with heavy polysomes, indicating decreased translational engagement in the absence of hnRNP U. This effect was particularly notable for transcripts encoding mitochondrial proteins, including AKAP1, whose reduced polysome association correlated with decreased protein expression and impaired colocalization at mitochondria. The data also indicate that mTOR signaling modulates RG4 dynamics and hnRNP U’s association with polysomes: pharmacologic inhibition of mTOR (e.g., rapamycin) increased RG4 folding and reduced hnRNP U binding to both RG4s and translating ribosomes, leading to decreased translation of RG4‑containing transcripts. Overall, the study establishes rG4s as a post-transcriptional layer of control essential for maintaining cellular energy balance. The study highlights the potential for small molecules (rG4 stabilizers) to act as precise metabolic modulators, opening new doors for treating metabolic and age-related diseases.
Learn more about EIRNABio’s polysome profiling services here.
Ribosome association inhibits stress-induced gene mRNA localization to stress granules
Genes and Development, 2025
Helton, N., Dodd, B., and Moon, S.

This study investigates how mRNA localization to stress granules (SGs) is regulated during cellular stress, with a focus on stress-induced transcripts. Using single-molecule imaging (smFISH), polysome profiling, and pharmacological perturbation of translation, the authors show that ribosome association limits mRNA partitioning into SGs. In particular, mRNAs associated with ribosomes are less likely to accumulate in SGs, whereas reducing ribosome association or promoting ribosome release increases SG localization for specific transcripts.
Upon arsenite treatment, polysome profiling showed global polysome collapse and translational repression. However, the uORF-containing stress-induced transcripts ATF4 and GADD34 remained translationally active (or became more ribosome-associated) and showed reduced SG localization relative to transcripts that were translationally repressed. The authors further demonstrated that inhibiting translation initiation (rocaglamide A) or releasing ribosomes (puromycin) increased ATF4 and GADD34 recruitment to SGs, while trapping ribosomes on mRNAs (harringtonine or emetine) reduced SG localization.
A key mechanistic finding is that even a single initiating ribosome can inhibit SG assembly and limit mRNA association with SGs. The authors also show that upstream open reading frames (uORFs) in ATF4 and GADD34 reporters reduce SG localization in a ribosome-dependent manner, supporting a model in which uORF-mediated ribosome association acts as a gatekeeper for RNA condensation during stress.
Overall, the key finding is that ribosome association acts as a gatekeeper for mRNA partitioning into stress granules. This highlights the integration of translational control and RNA localization in shaping cellular stress responses and influencing post-transcriptional gene regulation. Dysregulated SG dynamics are implicated in diseases like ALS, frontotemporal dementia, and cancer. Understanding that ribosome binding prevents SG entry could inform strategies to modulate SG composition or mRNA partitioning in pathological contexts.
Learn more about EIRNABio’s ribosome profiling and polysome profiling services here.



