Translatomics for lncRNA, bacteria, and pseudouridine
Recent Publications Harnessing the Power of Translatomics
Every week we provide a digest of a small number of recent interesting papers in the field of translatomics.
In this week’s Sunday papers,
- Li et al. reveal that LincNEAT1-derived NEAT1-31 micropeptide promotes macrophage phagocytosis by activating Aurora A–PI3K–AKT.
- Lee et al. show that multi-omics analysis of Streptomyces tsukubaensis shows coordinated transcriptional and translational control of FK506 biosynthesis.
- Liu et al. underline that programmable pseudouridine editing enables RNA codon expansion in mammalian cells.
LincNEAT1 Encoded‐NEAT1‐31 Promotes Phagocytosis by Directly Activating the Aurora‐A–PI3K–AKT Pathway
Advanced Science, 2025.
Li, J., Zhang, J., Li, X., Liu, X., Zeng, B., Luo, J., Wang, H., Zhang, H. and Gao, X.
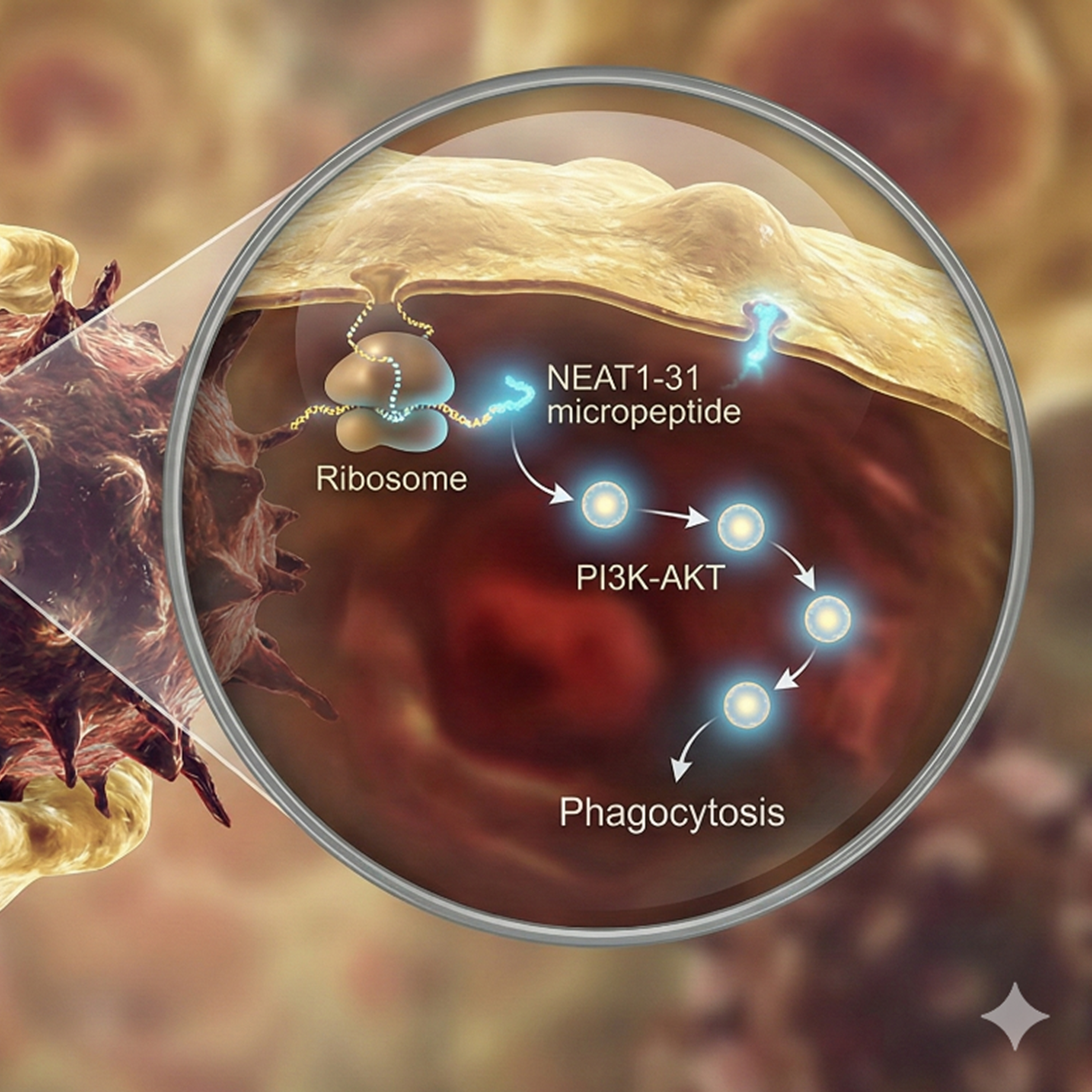
The study shows that the long non‑coding RNA LincNEAT1 encodes a previously unrecognized 31‑amino‑acid micropeptide, NEAT1‑31, which is highly upregulated in phagocytic macrophages and enhances their ability to engulf cancer cells. Using ribosome profiling (Ribo-seq), the authors identified NEAT1‑31 translation specifically in phagocytic macrophages. Purified NEAT1‑31 significantly increased macrophage phagocytosis of diverse cancer cell types both in vitro and in vivo and improved antigen presentation. It was found that NEAT1‑31 directly binds and activates Aurora‑A kinase, which in turn stimulates the PI3K–AKT signaling pathway, a molecular cascade that enhances phagocytosis in macrophages by promoting cytoskeletal changes and migration necessary for efficient phagocytosis. NEAT1‑31 also enhanced the efficacy of anti‑CD47 immunotherapy. CD47 overexpression on the surface of cancer cells facilitates immune detection evasion and anti-CD47 represents a potent class of checkpoint inhibitors designed to block this signal, thereby enabling macrophages to engulf and destroy the tumour cells. These findings reveal NEAT1‑31 as an intrinsic macrophage activator, uncovering a novel protein drug candidate that can bolster innate antitumor immunity and suggesting new therapeutic strategies to enhance cancer immunotherapy effectiveness.
Learn more about EIRNABio’s ribosome profiling services here.
Regulatory orchestration of FK506 biosynthesis in Streptomyces tsukubaensis NRRL 18488 revealed through systematic analysis
iScience, 2025.
Lee, N., Kim, W., Kim, J.H., Lee, Y., Hwang, S., Kim, G., Kim, H., Dan, Q., Schmidt, M., Yoon, Y.J., Cho, S., Palsson, B., Keasling, J.D., and Cho, B.-K.
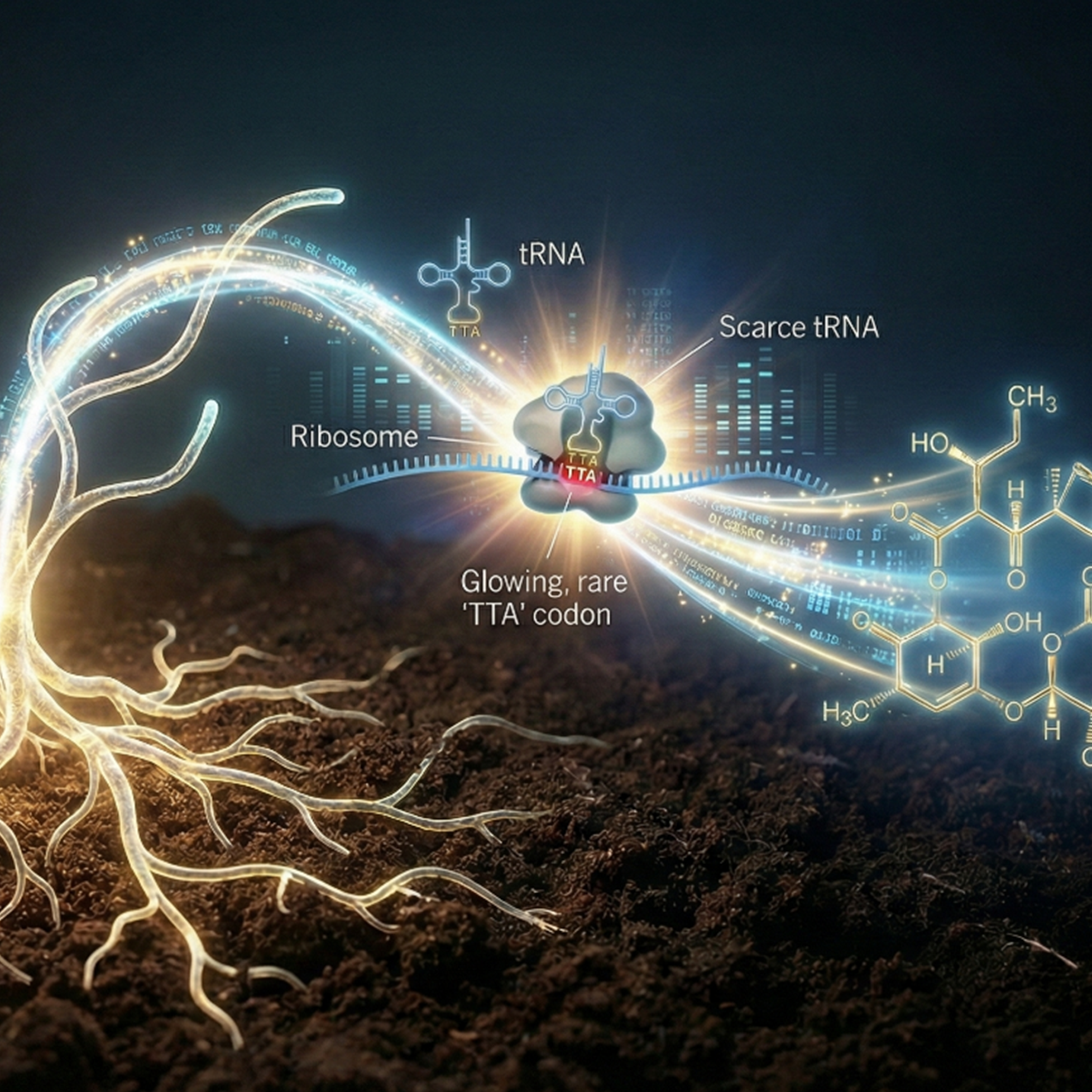
The authors performed an integrated multi‑omics analysis (genomics, transcriptomics, and translatomics) to decipher how Streptomyces tsukubaensis NRRL 18488 regulates biosynthesis of the immunosuppressant FK506 (tacrolimus). Streptomyces tsukubaensis NRRL 18488 is a soil-dwelling bacterium, isolated in 1984 from Tsukuba, Japan, famous as the producer of the Tacrolimus drug (FK506) which is used primarily to prevent organ rejection in transplant patients (kidney, liver, heart) by weakening the immune system. Completing and annotating the Streptomyces tsukubaensis 7.9 Mb genome enabled accurate characterization of the FK506 biosynthetic gene cluster (BGC). Transcriptome profiling during activation of FK506 production revealed a major shift from primary to secondary metabolism (where FK506 is produced) and highlighted coordinated upregulation of biosynthetic and precursor‑supply genes. Primary transcriptome mapping identified over 1,200 transcription units and indicated post‑transcriptional regulation of key precursor pathways, such as allylmalonyl‑CoA synthesis.
Ribosome profiling uncovered that AT‑rich codons, especially a rare leucine codon TTA, within the BGC, cause ribosome pausing and limit translational efficiency; substituting this codon alleviated pausing and boosted FK506 production. These findings illustrate complex regulation at transcriptional, post‑transcriptional, and translational levels that shapes secondary metabolite output. The work provides a systems‑level framework to guide strain engineering for enhanced natural product biosynthesis.
Learn more about EIRNABio’s ribosome profiling services here.
RNA codon expansion via programmable pseudouridine editing and decoding
Nature, 2025.
Liu, J., Yan, X., Wu, H., Ji, Z., Shan, Y., Wang, X., Ran, Y., Ma, Y., Li, C., Zhu, Y., Gu, R., Wen, H., Yi, C., and Chen, P.R.

In this work, the authors developed a novel RNA codon‑expansion (RCE) strategy that expands the genetic code in mammalian cells by using programmably introduced pseudouridine (Ψ) codons on specific mRNAs to encode non‑canonical amino acids (ncAAs). Traditional genetic code expansion methods reassign stop codons as “blank codons” which can interfere with normal translation termination resulting in off-target stop-codon readthrough at endogenous stop codons with subsequent proteome-wide toxicity. The RCE approach, however, installs bio‑orthogonal Ψ codons (ΨGA, ΨAA, ΨAG) into targeted transcripts via programmable guide RNAs and decodes them with engineered decoder tRNAs and aminoacyl‑tRNA synthetases, enabling precise ncAA incorporation. The authors used ribosome profiling to confirm specific decoding of the pseudouridine codons.
Ribosome profiling also confirmed the high specificity of RCE across the translatome and proteome, preserving endogenous stop codon usage, while the three Ψ‑codon–tRNA pairs were mutually orthogonal, allowing site‑specific and multiplexed ncAA incorporation in mammalian proteins. The system also functions compatibly with conventional genetic code expansion to permit dual ncAA incorporation within the same cell. This work opens a new approach for genetic alphabet expansion and programmable protein engineering in complex eukaryotic systems.
Learn more about EIRNABio’s ribosome profiling services here.



