Understanding Differential Gene Expression (DE) Analysis through Ribosome Profiling and RNA-seq
Introduction
In the dynamic landscape of molecular biology, differential gene expression (DE) analysis serves as a cornerstone for understanding the complex regulatory mechanisms underpinning biological processes and disease states. Two revolutionary techniques, ribosome profiling (Ribo-seq) and RNA sequencing (RNA-seq), have emerged as powerful tools for dissecting the nuances of gene expression at an unprecedented resolution. Integrating these methods offers a more comprehensive view of translational efficiency (TE) and gene expression dynamics. Here, we delve into the synergistic application of Ribo-seq and RNA-seq in DE analysis, highlighting how the EIRNA Bio Connect platform can expedite biological insights.
Ribosome Profiling: A Window into Translation Dynamics
Ribosome profiling provides a snapshot of active translation by sequencing ribosome-protected mRNA fragments1 allowing the identification of which mRNAs are being translated, thereby offering insights into the translational control of gene expression. Ribo-seq data are instrumental in revealing the intricacies of translation initiation, elongation, and termination, as well as uncovering hidden features such as alternative translation start sites and small open reading frames (sORFs).
RNA-seq: Quantifying the Transcriptome
Complementary to Ribo-seq, RNA-seq quantifies the transcriptome, providing a comprehensive view of the RNA species present within a cell at a given time. By sequencing the cDNA converted from RNA, researchers can measure the abundance of transcripts, uncover novel transcripts, and detect alternative splicing events2. This approach has improved our understanding of the complexity of eukaryotic transcriptomes.
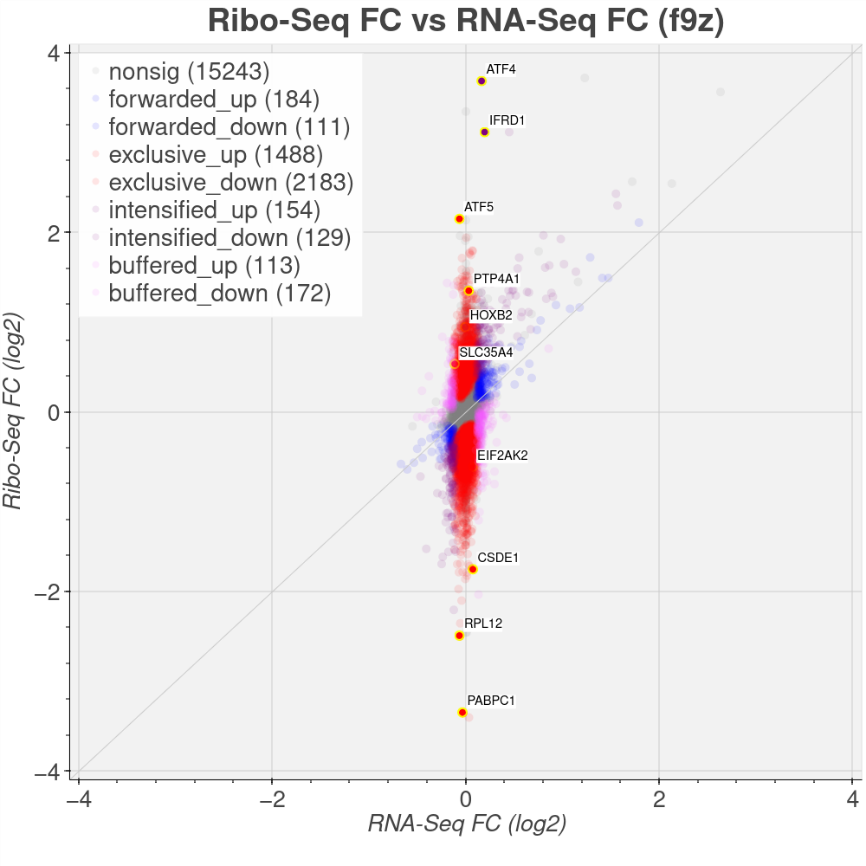
Figure 1 DeltaTE plot as implemented on the Connect platform where Ribo-seq and RNA-seq are combined for more complete DE analysis. Genes that are sensitive (downregulated) or resistant (upregulated) to stress induced by sodium arsenate treatment are easily discernible using the dynamic interactive Connect platform. Gene names can be dynamically highlighted and further explored (for illustration purposes a small selection of gene names are highlighted here). See also the first published study using Ribo-seq and RNA-seq to explore translational regulation upon EIF2 repression5.
The EIRNA Bio Connect Approach: Bridging Transcription and Translation
While RNA-seq and Ribo-seq individually offer valuable insights (Figure 1 volcano plot), combining these techniques unveils the full spectrum of gene expression regulation. The deltaTE approach3 which uses DESeq24 and implemented on EIRNA Bio Connect quantifies changes in TE between different conditions, accounting for changes in transcription. By comparing TE changes (deltaTE) across experimental conditions, genes whose translation is differentially regulated can be pinpointed, offering insights beyond mere changes in transcript levels – Figure 1.
Applications and Implications
The integration of Ribo-seq and RNA-seq for DE analysis can expand our understanding of cellular responses to various stimuli, disease mechanisms, and drug actions. Exemplified using EIRNA Bio’s high-quality data, Figure 1 shows how stress-responsive and stress-resistant genes that are translationally regulated can be identified. As translational control dominates the response to environmental signals, focusing on transcriptomics only could result in missing this important layer in gene expression regulation (Figure 2).

Challenges and Future Directions
Despite its immense potential, the combined application of Ribo-seq and RNA-seq poses computational and experimental challenges, including the normalization of data sets and the interpretation of complex changes in TE. EIRNA Bio Connect is a unique browser-based platform that enables combined Ribo-seq and RNA-seq DE analysis at the click of a button without the need for prior bioinformatic experience.
Conclusion
The confluence of Ribo-seq and RNA-seq represents a paradigm shift in DE analysis, moving beyond transcript abundance to a more nuanced understanding of translational regulation. As we continue to unravel the complexities of gene expression, integrating transcriptional and translational data will undoubtedly expand our understanding at an unparalleled depth.
Stay tuned for how EIRNA Bio Connect can provide actionable insights for your own data.
Over the course of the coming months, we will highlight additional EIRNA Bio Connect functionality to illustrate how our interactive platform can help advance your own research questions.
1. Ingolia NT, Ghaemmaghami S, Newman JR, & Weissman JS (2009). Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science, 324(5924), 218-223.
Wang Z, Gerstein M, & Snyder M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews Genetics, 10(1), 57-63.
Chothani S, Adami E, Ouyang JF, Viswanathan S, Hubner N, Cook SA, Schafer S, Rackham OJL (2019). deltaTE: Detection of Translationally Regulated Genes by Integrative Analysis of Ribo-seq and RNA-seq Data. Curr Protoc Mol Biol Dec;129(1):e108.
Love MI, Huber W, Anders S (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology, 15, 550.
Andreev DE, O’Connor PBF, Fahey C, Kenny EM, Terenin IM, Dmitriev SE, Cormican P, Morris DW, Shatsky IN, Baranov PV (2015) Translation of 5′ leaders is pervasive in genes resistant to eIF2 repression. Elife Jan 26:4:e03971.




